Complexity: A Guided Tour - Melanie Mitchell (2009)
Part IV. Network Thinking
Chapter 18. Evolution, Complexified
IN CHAPTER I I asked, “How did evolution produce creatures with such an enormous contrast between their individual simplicity and their collective sophistication?” Indeed, as illustrated by the examples we’ve seen in this book, the closer one looks at living systems, the more astonishing it seems that such intricate complexity could have been formed by the gradual accumulation of favorable mutations or the whims of historical accident. This very argument has been used from Darwin’s time to the present by believers in divine creation or other supernatural means of “intelligent design.”
The questions of how, why, and even if evolution creates complexity, and how complexity in biology might be characterized and measured, are still very much open. One of the most important contributions of complex systems research over the last few decades has been to demonstrate new ways to approach these age-old questions. In this chapter I describe some of the recent discoveries in genetics and the dynamics of genetic regulation that are giving us surprising new insights into the evolution of complex systems.
Genetics, Complexified
Often in science new technologies can open a floodgate of discoveries that change scientists’ views of a previously established field of study. We saw an example of this back in chapter 2—it was the invention of the electronic computer, and its capacity for modeling complex systems such as weather, that allowed for the demonstration of the existence of chaos. More recently, extremely powerful land and space-based telescopes have led to a flurry of discoveries in astronomy concerning so-called dark matter and dark energy, which seem to call into question much of what was previously accepted in cosmology.
No new set of technologies has had a more profound impact on an established field than the so-called molecular revolution in genetics over the last four decades. Technologies for rapidly copying, sequencing, synthesizing, and engineering DNA, for imaging molecular-level structures that had never been seen before, and for viewing expression patterns of thousands of different genes simultaneously; these are only a few examples of the feats of biotechnology in the late twentieth and early twenty-first centuries. And it seems that with each new technology allowing biologists to peer closer into the cell, more unexpected complexities appear.
At the time Watson and Crick discovered its structure, DNA was basically thought of as a string of genes, each of which coded for a particular protein that carried out some function in the cell. This string of genes was viewed essentially as the “computer program” of the cell, whose commands were translated and enacted by RNA, ribosomes, and the like, in order to synthesize the proteins that the genes stood for. Small random changes to the genome occurred when copying errors were made during the DNA duplication process; the long-term accumulation of those small random changes that happened to be favorable were the ultimate cause of adaptive change in biology and the origin of new species.
This conventional view has undergone monumental changes in the last 40 years. The term molecular revolution refers not only to the revolutionary new techniques in genetics, but also to the revolutionary new view of DNA, genes, and the nature of evolution that these techniques have provided.
What Is a Gene?
One casualty of the molecular revolution is the straightforward concept of gene. The mechanics of DNA that I sketched in chapter 6 still holds true—chromosomes contain stretches of DNA that are transcribed and translated to create proteins—but it turns out to be only part of the story. The following are a few examples that give the flavor of the many phenomena that have been and are being discovered; these phenomena are confounding the straightforward view of how genes and inheritance work.
· Genes are not like “beads on a string.” When I took high-school biology, genes and chromosomes were explained using the beads-on-a-string metaphor (and I think we even got to put together a model using pop-together plastic beads). However, it turns out that genes are not so discretely separated from one another. There are genes that overlap with other genes—i.e., they each code for a different protein, but they share DNA nucleotides. There are genes that are wholly contained inside other genes.
· Genes move around on their chromosome and between chromosomes. You may have heard of “jumping genes.” Indeed, genes can move around, rearranging the makeup of chromosomes. This can happen in any cell, including sperm and egg cells, meaning that the effects can be inherited. The result can be a much higher rate of mutation than comes from errors in DNA replication. Some scientists have proposed that these “mobile genetic elements” might be responsible for the differences observed between close relatives, and even between identical twins. The phenomenon of jumping genes has even been proposed as one of the mechanisms responsible for the diversity of life.
· A single gene can code for more than one protein. It had long been thought that there was a one-to-one correspondence between genes and proteins. A problem for this assumption arose when the human genome was sequenced, and it was discovered that while the number of different types of proteins encoded by genes may exceed 100,000, the human genome contains only about 25,000 genes. The recently discovered phenomena of alternative splicing and RNA editing help explain this discrepancy. These processes can alter messenger RNA in various ways after it has transcribed DNA but before it is translated into amino acids. This means that different transcription events of the same gene can produce different final proteins.
· In light of all these complications, even professional biologists don’t always agree on the definition of “gene.” Recently a group of science philosophers and biologists performed a survey in which 500 biologists were independently given certain unusual but real DNA sequences and asked whether each sequence qualified as a “gene,” and how confident they were of their answer. It turned out that for many of the sequences, opinion was split, with about 60% confident of one answer and 40% confident of the other answer. As stated in an article in Nature reporting on this work, “The more expert scientists become in molecular genetics, the less easy it is to be sure about what, if anything, a gene actually is.”
· The complexity of living systems is largely due to networks of genes rather than the sum of independent effects of individual genes. As I described in chapter 16, genetic regulatory networks are currently a major focus of the field of genetics. In the old genes-as-beads-on-a-string view, as in Mendel’s laws, genes are linear—each gene independently contributes to the entire phenotype. The new, generally accepted view, is that genes in a cell operate in nonlinear information-processing networks, in which some genes control the actions of other genes in response to changes in the cell’s state—that is, genes do not operate independently.
· There are heritable changes in the function of genes that can occur without any modification of the gene’s DNA sequence. Such changes are studied in the growing field of epigenetics. One example is so-called DNA methylation, in which an enzyme in a cell attaches particular molecules to some parts of a DNA sequence, effectively “turning off” those parts. When this occurs in a cell, all descendents of that cell will have the same DNA methylation. Thus if DNA methylation occurs in a sperm or egg cell, it will be inherited.
On the one hand, this kind of epigenetic effect happens all the time in our cells, and is essential for life in many respects, turning off genes that are no longer needed (e.g., once we reach adulthood, we no longer need to grow and develop like a child; thus genes controlling juvenile development are methylated). On the other hand, incorrect or absent methylation is the cause of some genetic disorders and diseases. In fact, the absence of necessary methylation during embryo development is thought by some to be the reason so many cloned embryos do not survive to birth, or why so many cloned animals that do survive have serious, often fatal disorders.
· It has recently been discovered that in most organisms a large proportion of the DNA that is transcribed by RNA is not subsequently translated into proteins. This so-called noncoding RNA can have many regulatory effects on genes, as well as functional roles in cells, both of which jobs were previously thought to be the sole purview of proteins. The significance of non-coding RNAs is currently a very active research topic in genetics.
Genetics has become very complicated indeed. And the implications of all these complications for biology are enormous. In 2003 the Human Genome Project published the entire human genome—that is, the complete sequence of human DNA. Although a tremendous amount was learned from this project, it was less than some had hoped. Some had believed that a complete mapping of human genes would provide a nearly complete understanding of how genetics worked, which genes were responsible for which traits, and that this would guide the way for revolutionary medical discoveries and targeted gene therapies. Although there have been several discoveries of certain genes that are implicated in particular diseases, it has turned out that simply knowing the sequence of DNA is not nearly enough to understand a person’s (or any complex organism’s) unique collection of traits and defects.
One sector that pinned high hopes on the sequencing of genes is the international biotechnology industry. A recent New York Times article reported on the effects that all this newly discovered genetic complexity was having on biotech: “The presumption that genes operate independently has been institutionalized since 1976, when the first biotech company was founded. In fact, it is the economic and regulatory foundation on which the entire biotechnology industry is built.”
The problem is not just that the science underlying genetics is being rapidly revised. A major issue lurking for biotech is the status of gene patents. For decades biotech companies have been patenting particular sequences of human DNA that were believed to “encode a specific functional product.” But as we have seen above, many, if not most, complex traits are not determined by the exact DNA sequence of a particular gene. So are these patents defensible? What if the “functional product” is the result of epigenetic processes acting on the gene or its regulators? Or what if the product requires not only the patented gene but also the genes that regulate it, and the genes that regulate those genes, and so on? And what if those regulatory genes are patented by someone else? Once we leave the world of linear genes and encounter essential nonlinearity, the meaning of these patents becomes very murky and may guarantee the employment of patent lawyers and judges for a long time to come. And patents aren’t the only problem. As the New York Times pointed out, “Evidence of a networked genome shatters the scientific basis for virtually every official risk assessment of today’s commercial biotech products, from genetically engineered crops to pharmaceuticals.”
Not only genetics, but evolutionary theory as a whole has been profoundly challenged by these new genetic discoveries. A prominent example of this is the field of “Evo-Devo.”
Evo-Devo
Evo-Devo is the nickname for “evolutionary developmental biology.” Many people are very excited about this field and its recent discoveries, which are claimed to explain at least three big mysteries of genetics and evolution: (1) Humans have only about 25,000 genes. What is responsible for our complexity? (2) Genetically, humans are very similar to many other species. For example, more than 90% of our DNA is shared with mice and more than 95% with chimps. Why are our bodies so different from those of other animals? (3) Supposing that Stephen Jay Gould and others are correct about punctuated equilibria in evolution, how could big changes in body morphology happen in short periods of evolutionary time?
It has recently been proposed that the answer to these questions lies, at least in part, in the discovery of genetic switches.
The fields of developmental biology and embryology study the processes by which a fertilized single egg cell becomes a viable multibillion-celled living organism. However, the Modern Synthesis’s concern was with genes; in the words of developmental biologist Sean Carroll, it treated developmental biology and embryology as a “ ‘black box’ that somehow transformed genetic information into three-dimensional, functional animals.” This was in part due to the view that the huge diversity of animal morphology would eventually be explained by large differences in the number of and DNA makeup of genes.
In the 1980s and 1990s, this view became widely challenged. As I noted above, DNA sequencing had revealed the extensive similarities in DNA among many different species. Advances in genetics also produced a detailed understanding of the mechanisms of gene expression in cells during embryonic and fetal development. These mechanisms turned out to be quite different from what was generally expected. Embryologists discovered that, in all complex animals under study, there is a small set of “master genes” that regulate the formation and morphology of many of the animal’s body parts. Even more surprising, these master genes were found to share many of the same sequences of DNA across many species with extreme morphological differences, ranging from fruit flies to humans.
Given that their developmental processes are governed by the same genes, how is it that these different animals develop such different body parts? Proponents of Evo-Devo propose that morphological diversity among species is, for the most part, not due to differences in genes but in genetic switches that are used to turn genes on and off. These switches are sequences of DNA—often several hundred base pairs in length—that do not code for any protein. Rather they are part of what used to be called “junk DNA,” but now have been found to be used in gene regulation.
Figure 18.1 illustrates how switches work. A switch is a sequence of non-coding DNA that resides nearby a particular gene. This sequence of molecules typically contains on the order of a dozen signature subsequences, each of which chemically binds with a particular protein, that is, the protein attaches to the DNA string. Whether or not the nearby gene gets transcribed, and how quickly, depends on the combination of proteins attached to these subsequences. Proteins that allow transcription create strong binding sites for RNA molecules that will do the transcribing; proteins that prevent transcription block these same RNA molecules from binding to the DNA. Some of these proteins can negate the effects of others.
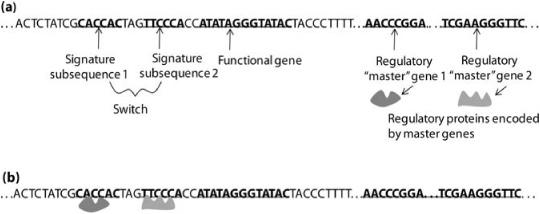

FIGURE 18.1. Illustration of genetic “switches.” (a) A DNA sequence, containing a switch with two signature subsequences, a functional gene turned on by that switch, and two regulatory master genes. The regulatory master genes give rise to regulatory proteins. (b) The regulatory proteins bind to the signature subsequences, switching on the functional gene—that is, allowing it to be transcribed.
Where do these special regulator proteins come from? Like all proteins, they come from genes, in this case regulatory genes that encode such proteins in order to turn other genes on or off, depending on the current state of the cell. How do these regulatory genes determine the current state of the cell? By the presence or absence of proteins that signal the state of the cell by binding to the regulatory genes’ own switches. Such proteins are often encoded by other regulatory genes, and so forth.
In summary, genetic regulatory networks are made up of several different kinds of entities, including functional genes that encode proteins (and sometimes noncoding RNA) for cellular maintenance or building, and regulatorygenes that encode proteins (and sometimes noncoding RNA) that turn other genes on or off by binding to DNA “switches” near to the gene in question.
I can now give Evo-Devo’s answers to the three questions posed at the beginning of this section. Humans (and other animals) can be more complex than their number of genes would suggest for many reasons, some listed above in the “What Is a Gene” section. But a primary reason is that genetic regulatory networks allow a huge number of possibilities for gene expression patterns, since there are so many possible ways in which proteins can be attached to switches.
The reason we humans can share so many genes with other creatures quite different from us is that, although the genes might be the same, the sequences making up switches have often evolved to be different. Small changes in switches can produce very different patterns of genes turning on and off during development. Thus, according to Evo-Devo, the diversity of organisms is largely due to evolutionary modifications of switches, rather than genes. This is also the reason that large changes in morphology—possibly including speciation—can happen swiftly in evolutionary time: the master genes remain the same, but switches are modified. According to Evo-Devo, such modifications—in the parts of DNA long thought of as “junk”—are the major force in evolution, rather than the appearance of new genes. Biologist John Mattick goes so far as to say, “The irony … is that what was dismissed as junk [DNA] because it wasn’t understood will turn out to hold the secret of human complexity.”
One striking instance of Evo-Devo in action is the famous example of the evolution of finches’ beaks. As I described in chapter 5, Darwin observed large variations in beak size and shape among finches native to the Galápagos Islands. Until recently, most evolutionary biologists would have assumed that such variations resulted from a gradual process in which chance mutations of several different genes accumulated. But recently, a gene called BMP4 was discovered that helps control beak size and shape by regulating other genes that produce bones. The more strongly BMP4 is expressed during the birds’ development, the larger and stronger their beaks. A second gene, called calmodulin, was discovered to be associated with long, thin beaks. As Carol Kaesuk Yoon reported in the New York Times, “To verify that the BMP4 gene itself could indeed trigger the growth of grander, bigger, nut-crushing beaks, researchers artificially cranked up the production of BMP4 in the developing beaks of chicken embryos. The chicks began growing wider, taller, more robust beaks similar to those of a nut-cracking finch …. As with BMP4, the more that calmodulin was expressed, the longer the beak became. When scientists artificially increased calmodulin in chicken embryos, the chicks began growing extended beaks, just like a cactus driller …. So, with just these two genes, not tens or hundreds, the scientists found the potential to re-create beaks, massive or stubby or elongated.” The conclusion is that large changes in the morphology of beaks (and other traits) can take place rapidly without the necessity of waiting for many chance mutations over a long period of time.
Another example where Evo-Devo is challenging long-held views about evolution concerns the notion of convergent evolution. In my high school biology class, we learned that the octopus eye and the human eye—greatly different in morphology—were examples of convergent evolution: eyes in these two species evolved completely independently of one another as a consequence of natural selection acting in two different environments in which eyes were a useful adaptation.
However, recent evidence has indicated that the evolution of these two eyes was not as independent as previously thought. Humans, octopi, flies, and many other species have a common gene called PAX6, which helps direct the development of eyes. In a strange but revealing experiment, the Swiss biologist Walter Gehring took PAX6 genes from mice and inserted them into the genomes of fruit flies. In particular, in different studies, PAX6 was inserted in three different parts of the genome: those that direct the development of legs, wings, and antennae, respectively. The researchers got eerie results: eye-like structures formed on flies’ legs, wings, and antennae. Moreover, the structures were like fly eyes, not mouse eyes. Gehring’s conclusion: the eye evolved not many times independently, but only once, in a common ancestor with the PAX6 gene. This conclusion is still quite controversial among evolutionary biologists.
Although genetic regulatory networks directed by master genes can produce great diversity, they also enforce certain constraints on evolution. Evo-Devo scientists claim that the types of body morphology (called body plans) any organism can have are highly constrained by the master genes, and that is why only a few basic body plans are seen in nature. It’s possible that genomes vastly different from ours could result in new types of body plans, but in practice, evolution can’t get us there because we are so reliant on the existing regulatory genes. Our possibilities for evolution are constrained. According to Evo-Devo, the notion that “every trait can vary indefinitely” is wrong.
Genetic Regulation and Kauffman’s “Origins of Order”
Stuart Kauffman is a theoretical biologist who has been thinking about genetic regulatory networks and their role in constraining evolution for over forty years, long before the ascendency of Evo-Devo. He has also thought about the implications for evolution of the “order” we see emerging from such complex networks.
Kauffman is a legendary figure in complex systems. My first encounter with him was at a conference I attended during my last year of graduate school. His talk was the very first one at the conference, and I must say that, for me at the time, it was the most inspiring talk I had ever heard. I don’t remember the exact topic; I just remember the feeling I had while listening that what he was saying was profound, the questions he was addressing were the most important ones, and I wanted to work on this stuff too.
Kauffman started his career with a short stint as a physician but soon moved to genetics research. His work was original and influential; it earned him many academic accolades, including a MacArthur “genius” award, as well as a faculty position at the Santa Fe Institute. At SFI seminars, Kauffman would sometimes chime in from the audience with, “I know I’m just a simple country doctor, but … ” and would spend a good five minutes or more fluently and eloquently giving his extemporaneous opinion on some highly technical topic that he had never thought about before. One science journalist called him a “world-class intellectual riffer,” which is an apt description that I interpret as wholly complimentary.

Stuart Kauffman (Photograph by Daryl Black, reprinted with permission.)
Stuart’s “simple country doctor” humble affect belies his personality. Kauffman is one of Complex Systems’ big thinkers, a visionary, and not what you would call a “modest” or “humble” person. A joke at SFI was that Stuart had “patented Darwinian evolution,” and indeed, he holds a patent on techniques for evolving protein sequences in the laboratory for the purpose of discovering new useful drugs.
RANDOM BOOLEAN NETWORKS
Kauffman was perhaps the first person to invent and study simplified computer models of genetic regulatory networks. His model was a structure called a Random Boolean Network (RBN), which is an extension of cellular automata. Like any network, an RBN consists of a set of nodes and links between the nodes. Like a cellular automaton, an RBN updates its nodes’ states in discrete time steps. At each time step each node can be in either state on or state off.
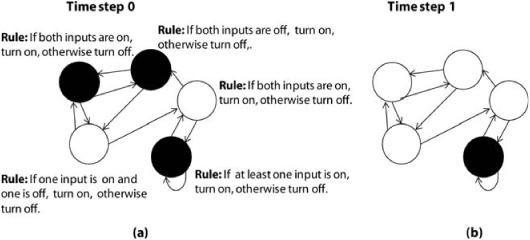
FIGURE 18.2. (a) A random Boolean network with five nodes. The in-degree (K) of each node is equal to 2. At time step 0, each node is in a random initial state: on (black) or off (white). (b) Time step 1 shows the network after each node has updated its state.
The property that on and off are the only allowed states is where the term Boolean comes in: a Boolean rule (or function) is one that gets some number of inputs, each equal to either 0 or 1, and from those inputs it produces an output of a 0 or 1. Such rules are named after the mathematician George Boole, who did extensive mathematical research on them.
In an RBN, links are directional: if node A links to node B, node B does not necessarily (but can possibly) link to node A. The in-degree of each node (the number of links from other nodes to that node) is the same for each node—let’s call that number K.
Here is how to build an RBN: for each node, create in-links to that node from K other randomly chosen nodes (including, possibly, a self-link), and give that node a Boolean rule, chosen randomly, that inputs K on or off states and outputs a single on or off state (figure 18.2a).
To run the RBN, give each node an initial state of on or off chosen at random. Then at each time step, each node transmits its state to the nodes it links to, and receives as input the states from the nodes that link to it. Each node then applies its rule to its input to determine its state at the next time step. All this is illustrated in figure 18.2, which shows the action of an RBN of five nodes, each with two inputs, over one time step.
RBNs are similar to cellular automata, but with two major differences: nodes are connected not to spatially neighboring nodes but at random, and rather than all nodes having an identical rule, each node has its own rule.
In Kauffman’s work, the RBN as a whole is an idealized model of a genetic regulatory network, in which “genes” are represented by nodes, and “gene A regulates gene B” is represented by node A linking to node B. The model is of course vastly simpler than real genetic networks. Using such idealized models in biology is now becoming common, but when Kauffman started this work in the 1960s, it was less well accepted.
LIFE AT THE EDGE OF CHAOS
Kauffman and his students and collaborators have done a raft of simulations of RBNs with different values of the in-degree K for each node. Starting from a random initial state, and iterated over a series of time steps, the nodes in the RBN change state in random ways for a while, and finally settle down to either a fixed point (all nodes’ states remain fixed) or an oscillation (the state of the whole network oscillates with some small period), or do not settle down at all, with random-looking behavior continuing over a large number of iterations. Such behavior is chaotic, in that the precise trajectory of states of the network have sensitive dependence on the initial state of the network.
Kauffman found that the typical final behavior is determined by both the number of nodes in the network and each node’s in-degree K. As K is increased from 1 (i.e., each node has exactly one input) all the way up to the total number of nodes (i.e., each node gets input from all other nodes, including itself), the typical behavior of the RBNs moves through the three different “regimes” of behavior (fixed-point, oscillating, chaotic). You might notice that this parallels the behavior of the logistic map as R is increased (cf. chapter 2). At K = 2 Kauffman found an “interesting” regime—neither fixed point, oscillating, or completely chaotic. In analogy with the term “onset of chaos” used with the logistic map, he called this regime the “edge of chaos.”
Assuming the behavior of his RBNs reflected the behavior of real genetic networks, and making an analogy with the phases of water as temperature changes, he concluded that “the genomic networks that control developmentfrom zygote to adult can exist in three major regimes: a frozen ordered regime, a gaseous chaotic regime, and a kind of liquid regime located in the region between order and chaos.”
Kauffman reasoned that, for an organism to be both alive and stable, the genetic networks his RBNs modeled had to be in the interesting “liquid” regime—not too rigid or “frozen,” and not too chaotic or “gaseous.” In his own words, “life exists at the edge of chaos.”
Kauffman used the vocabulary of dynamical systems theory—attractors, bifurcations, chaos—to describe his findings. Suppose we call a possible configuration of the nodes’ states a global state of the network. Since RBNs have a finite number of nodes, there are only a finite number of possible global states, so if the network is iterated for long enough it will repeat one of the global states it has already been in, and hence cycle through the next series of states until it repeats that global state again. Kauffman called this cycle an “attractor” of the network. By performing many simulations of RBNs, he estimated that the average number of different attractors produced in different networks with K = 2 was approximately equal to the square root of the number of nodes.
Next came a big leap in Kauffman’s interpretation of this model. Every cell in the body has more or less identical DNA. However, the body has different types of cells: skin cells, liver cells, and so forth. Kauffman asserted that what determines a particular cell type is the pattern of gene expression in the cell over time—I have described above how gene expression patterns can be quite different in different cells. In the RBN model, an attractor, as defined above, is a pattern over time of “gene expression.” Thus Kauffman proposed that an attractor in his network represents a cell type in an organism’s body. Kauffman’s model thus predicted that for an organism with 100,000 genes, the number of different cell types would be approximately the square root of 100,000, or 316. This is not too far from the actual number of cell types identified in humans—somewhere around 256.
At the time Kauffman was doing these calculations, it was generally believed that the human genome contained about 100,000 genes (since the human body uses about 100,000 types of proteins). Kauffman was thrilled that his model had come close to correctly predicting the number of cell types in humans. Now we know that the human genome contains only about 25,000 genes, so Kauffman’s model would predict about 158 cell types.
THE ORIGIN OF ORDER
The model wasn’t perfect, but Kauffman believed it illustrated his most important general point about living systems: that natural selection is in principle not necessary to create a complex creature. Many RBNs with K = 2 exhibited what he termed “complex” behavior, and no natural selection or evolutionary algorithm was involved. His view was that once a network structure becomes sufficiently complex—that is, has a large number of nodes controlling other nodes—complex and “self-organized” behavior will emerge. He says,
Most biologists, heritors of the Darwinian tradition, suppose that the order of ontogeny is due to the grinding away of a molecular Rube Goldberg machine, slapped together piece by piece by evolution. I present a countering thesis: most of the beautiful order seen in ontogeny is spontaneous, a natural expression of the stunning self-organization that abounds in very complex regulatory networks. We appear to have been profoundly wrong. Order, vast and generative, arises naturally.
Kauffman was deeply influenced by the framework of statistical mechanics, which I described in chapter 3. Recall that statistical mechanics explains how properties such as temperature arise from the statistics of huge numbers of molecules. That is, one can predict the behavior of a system’s temperature without having to follow the Newtonian trajectory of every molecule. Kauffman similarly proposed that he had found a statistical mechanics law governing the emergence of complexity from huge numbers of interconnected, mutually regulating components. He termed this law a “candidate fourth law of thermodynamics.” Just as the second law states that the universe has an innate tendency toward increasing entropy, Kauffman’s “fourth law” proposes that life has an innate tendency to become more complex, which is independent of any tendency of natural selection. This idea is discussed at length in Kauffman’s book, The Origins of Order. In Kauffman’s view, the evolution of complex organisms is due in part to this self-organization and in part to natural selection, and perhaps self-organization is really what predominates, severely limiting the possibilities for selection to act on.
Reactions to Kauffman’s Work
Given that Kauffman’s work implies “a fundamental reinterpretation of the place of selection in evolutionary theory,” you can imagine that people react rather strongly to it. There are a lot of huge fans of this work (“His approach opens up new vistas”; it is “the first serious attempt to model a complete biology”). On the other side, many people are highly skeptical of both his results and his broad interpretations of them. One reviewer called Kauffman’s writing style “dangerously seductive” and said of The Origins of Order, “There are times when the bracing walk through hyperspace seems unfazed by the nagging demands of reality.”
Indeed, the experimental evidence concerning Kauffman’s claims is not all on his side. Kauffman himself admits that regarding RBNs as models of genetic regulatory networks requires many unrealistic assumptions: each node can be in only one of two states (whereas gene expression has different degrees of strength), each has an identical number of nodes that regulate it, and all nodes are updated in synchrony at discrete time steps. These simplifications may ignore important details of genetic activity.
Most troublesome for his theory are the effects of “noise”—errors and other sources of nondeterministic behavior—that are inevitable in real-world complex systems, including genetic regulation. Biological genetic networks make errors all the time, yet they are resilient—most often our health is not affected by these errors. However, simulations have shown that noise has a significant effect on the behavior of RBNs, and sometimes will prevent RBNs from reaching a stable attractor. Even some of the claims Kauffman made specifically about his RBN results are not holding up to further scrutiny. For example, recall Kauffman’s claim that the number of attractors that occur in a typical network is close to the square root of the number of nodes, and his interpretation of this fact in terms of cell-types. Additional simulations have shown that the number of attractors is actually not well approximated by the square root of the number of nodes. Of course this doesn’t necessarily mean that Kauffman is wrong in his broader claims; it just shows that there is considerably more work to be done on developing more accurate models. Developing accurate models of genetic regulatory networks is currently a very active research area in biology.
Summary
Evolutionary biology is still working on answering its most important question: How does complexity in living systems come about through evolution? As we have seen in this chapter, the degree of complexity in biology is only beginning to be fully appreciated. We also have seen that many major steps are being taken toward understanding the evolution of complexity. One step has been the development of what some have called an “extended Synthesis,” in which natural selection still plays an important role, but other forces—historical accidents, developmental constraints, and self-organization—are joining natural selection as explanatory tools. Evolutionists, particularly in the United States, have been under attack from religious extremists and are often on the defensive, reluctant to admit that natural selection may not be the entire story. As biologists Guy Hoelzer, John Pepper, and Eric Smith have written about this predicament: “It has essentially become a matter of social responsibility for evolutionary biologists to join the battle in defense of Darwinism, but there is a scientific cost associated with this cultural norm. Alternative ways of describing evolutionary processes, complementary to natural selection, can elicit the same defensive posture without critical analysis.”
Evolutionary biologist Dan McShea has given me a useful way to think about these various issues. He classifies evolutionists into three categories: adaptationists, who believe that natural selection is primary; historicists, who give credit to historical accident for many evolutionary changes; and structuralists, such as Kauffman, who focus on how organized structure comes about even in the absence of natural selection. Evolutionary theory will be unified only when these three groups are able to show how their favored forces work as an integrated whole.
Dan also gave me an optimistic perspective on this prospect: “Evolutionary biology is in a state of intellectual chaos. But it’s an intellectual chaos of a very productive kind.”